Course description
This two-half-day training course introduces participants to genomic epidemiology and phylogenetic analysis of Brucella spp. using a realistic One Health scenario, and it explores the application of combined genomic and epidemiological data for outbreak investigations. Through guided analysis of Brucella spp. genomes, participants will evaluate the genomic diversity within the genus and gain practical skills for interpreting results from taxonomic analysis and typing, reading phylogenetic trees and assessing outbreak clusters. Importantly, participants will integrate metadata and epidemiological data with the genomic results, to infer outbreak sources and transmission dynamics.
The course includes a simulation exercise to reinforce practical understanding.
Outline
Target Audience
Microbiologists, clinicians and epidemiologists with basic bioinformatics skills
Pre-requisites: Equipment and Software requirements
To participate effectively in this course, ensure you have access to the necessary materials and that your machine meets the following requirements and has the appropriate tools installed:
- A laptop with internet access.
- Sample datasets (provided via ScienceData.dk: https://sciencedata.dk/shared/e9cba091ad1d946970bfdd67e8491b2a).
- For command-line usage: macOS or Linux recommended; Windows users should use WSL (Windows Subsystem for Linux) or Docker.
- Mamba or Conda for managing software environments and packages (installed through Miniforge for for example: https://github.com/conda-forge/miniforge)
Participants may choose to use web-based (basic) tools or command-line (advanced) tools depending on their comfort and experience:
- Pre-install the command-line tools before Session 1 for a smooth workflow.
- Web tools require no installation (access via provided links).
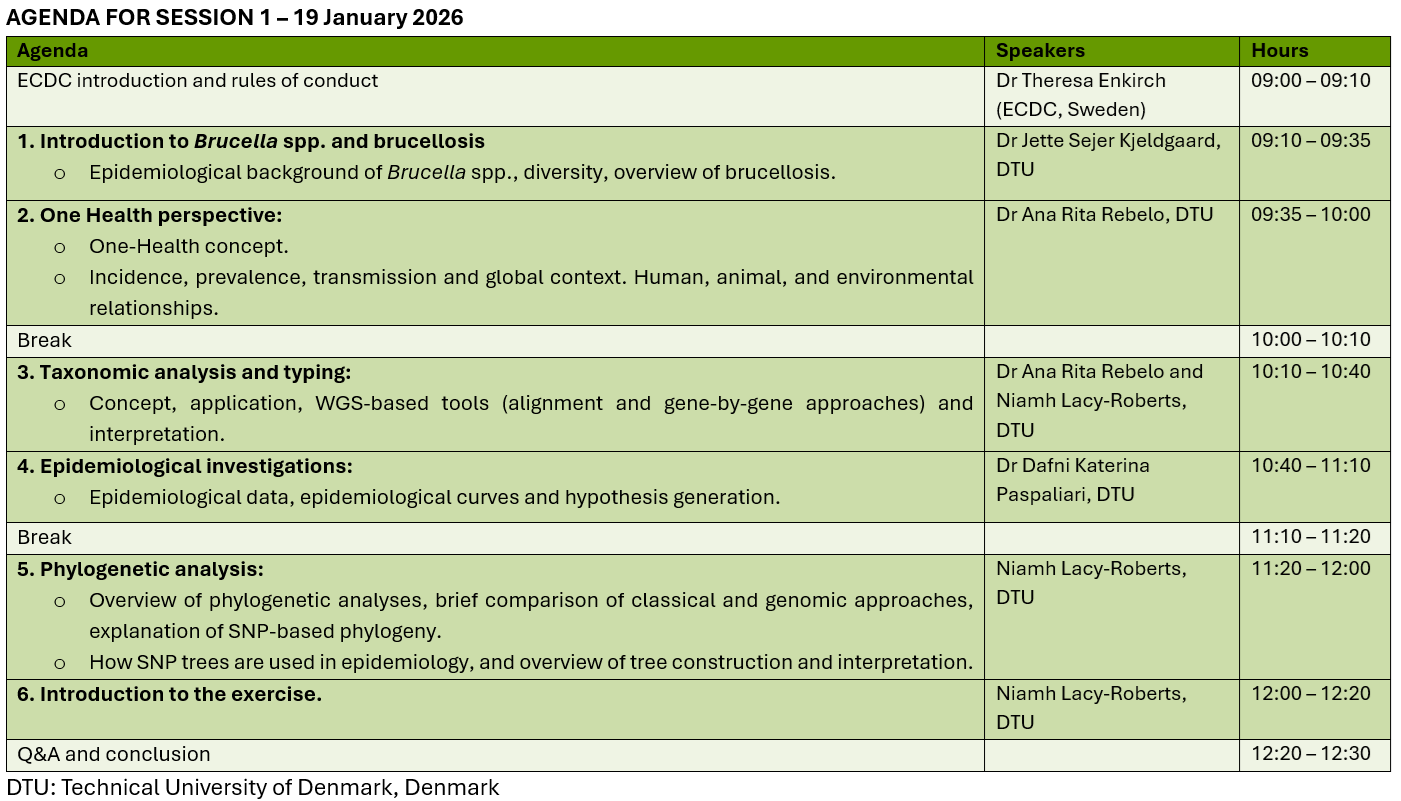
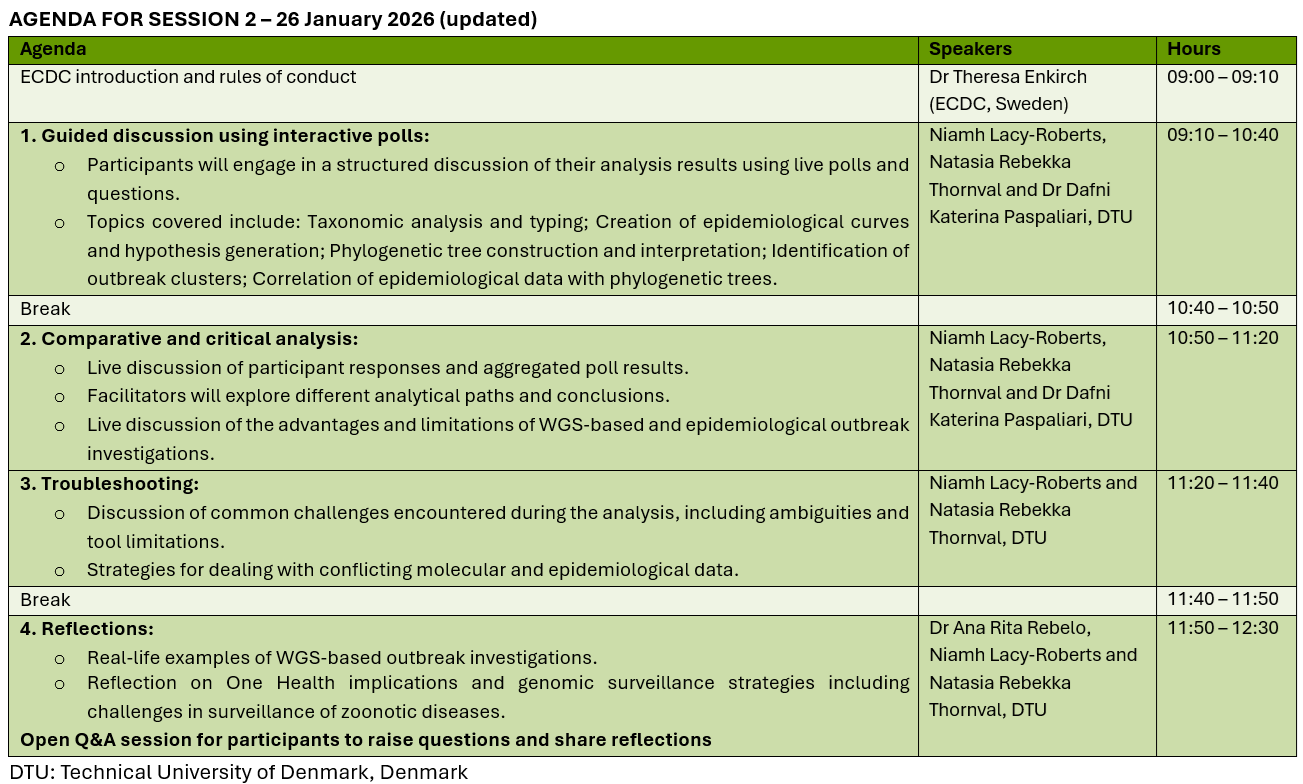
Curriculum and session details
The training is two half-days (Monday 19 January and Monday 26 January). Each session lasts 3.5 hours, from 9:00 to 12:30.
The practical exercises assigned in Session 1 should be completed before the next session. The discussion of the simulated exercise will take place during Session 2.
Suggested Readings
- Centre for Disease Prevention and Control (CDC). About Brucellosis. CDC, 2025. Available at: https://www.cdc.gov/brucellosis/about/index.html
- Centre for Disease Prevention and Control (CDC). Clinical Overview of Brucellosis. CDC, 2025. Available at: https://www.cdc.gov/brucellosis/hcp/clinical-overview/index.html
- World Health Organization (WHO). Brucellosis. WHO, 2025. Available at: https://www.who.int/news-room/fact-sheets/detail/brucellosis
- Whatmore AM, Perrett LL, MacMillan AP. Characterisation of the genetic diversity of Brucella by multilocus sequencing. BMC Microbiol. 2007;7:34. doi: 10.1186/1471-2180-7-34.
- Whatmore AM, Koylass MS, Muchowski J, et al. Extended Multilocus Sequence Analysis to Describe the Global Population Structure of the Genus Brucella: Phylogeography and Relationship to Biovars. Front Microbiol. 2016;7:2049. doi: 10.3389/fmicb.2016.02049.
- Whatmore AM, Foster JT. Emerging diversity and ongoing expansion of the genus Brucella. Infect Genet Evol. 2021;92:104865. doi: 10.1016/j.meegid.2021.104865.