Course description
This two-half-day training course introduces participants to the major pathotypes of the diverse pathogen E. coli, and to the genetic virulence factors that characterise them. Differentiating among gastrointestinal (GI) E. coli pathotypes can be challenging, particularly when relying on phenotypic or conventional laboratory methods. Currently, many clinical laboratories use PCR panels that can confirm a range of pathogenic E. coli and additional GI bacterial pathogens in a single run, but provide no additional typing. In most clinical laboratories, there is still a need for an isolate to perform additional typing and further cluster analysis. Genomic analyses enable the detection of virulence genes and other pathotype-specific markers, as well as the application of other typing and clustering methods. There are, however, genomic challenges in distinguishing Shigella from Enteroinvasive E. coli (EIEC), which will also be addressed. The course will explore the application of virulence gene profiling to predict pathotypes and, hence, the pathogenicity of E. coli isolates using a combination of traditional typing methods and virulence profiling. Through guided analysis of E. coli genomes, participants will evaluate different virulence identification tools for E. coli in combination with subtyping methods and use them to interpret the pathogenic potential of E. coli isolates. Participants will go through a critical review of E. coli sequence QC data to evaluate potential outlying results, including intra-species contamination, and will then be introduced to online or graphical user-interface-based tools for typing and virulence profiling.
The course includes an exercise on virulence profiling to reinforce practical understanding.
Target Audience
Microbiologists, clinicians, and epidemiologists with basic bioinformatics skills
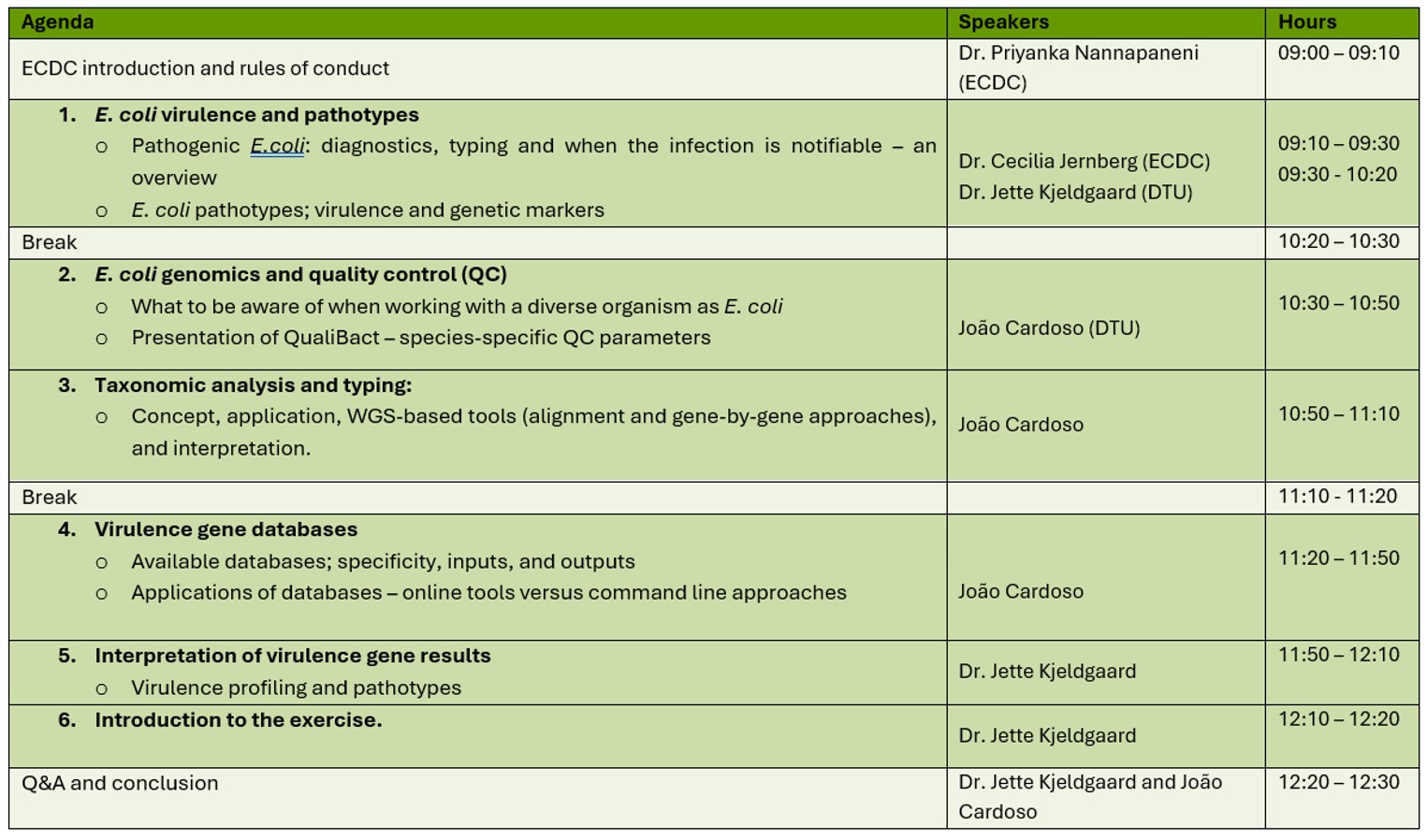
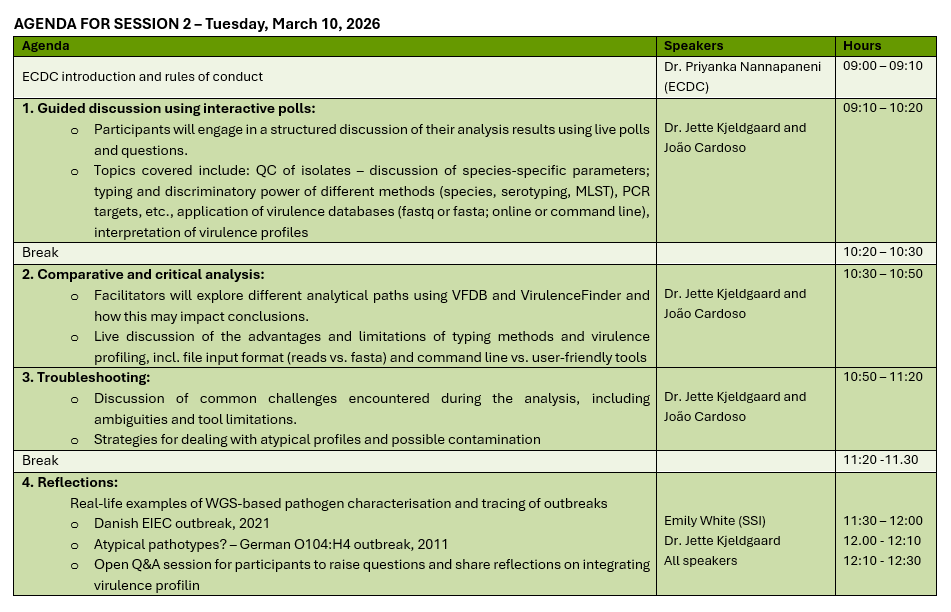 March 10, 2026
March 10, 2026